Making an Antibody
The question of the day - can I make an antibody? And even more interesting - can I make a good antibody?
Can I make an antibody?
I had a day off today. I’ve been reading a book about the immune system, and then I came across this article at breakfast: How to Design Antibodies - Asimov Press. The article offers a step by step guide to making an antiboddy, and also mentions an antibody design contest run by www.adaptyvbio.com/. I’ve been wanting to do something like this (well more specifically something like a CACHE challenge to attempt to make a drug, but that seems expensive). And thought it would be fun to take a crack at it. I had big goals initially and then decided I’d refocus on this more achievable goal for the day: make an antibody.
My immune system does it all the time, how hard could it be?
Target of interest: RBX1
The latest iteration of the antibody design competition focuses on RBX1. Here’s what they say about it:
The protein combines an intrinsically disordered N-terminal region with a stabilized C-terminal RING-H2 finger domain coordinating three zinc ions, a dual structural challenge for binder design. Disrupting RBX1 function could inhibit CRL-mediated protein degradation, with applications in cancer therapy where CRL activity is frequently dysregulated.
To understand a little more about why we’re targetting RBX1, it was useful to learn that:
The ubiquitination pathway relies on the sequential action of three enzymes: an E1-activating enzyme, an E2-conjugating enzyme, and an E3 ligase. Cullin RING E3 ubiquitin ligases (CRLs) constitute the major subfamily of E3 ligases that catalyze the transfer of ubiquitin from the E2-conjugating enzyme to the target substrate. source
And with a little googling, I’ve learned that ubiquitination is the process of adding ubiquitin to target substrate proteins. Which….. seems to do many things. But among those is protein degradation - tagging proteins for reycling - which brings us back to our antibody challenge, where they state that CRL-mediated protein degradataion is frequently dysregulated in cancer.
a sidequest on PROTACs
Related but probably unrelated learning along the way: PROTACs are a family of drugs which use ubiquitination to degrade specific proteins. Here’s what wikipedia says:
A proteolysis targeting chimera (PROTAC) is a molecule that can remove specific unwanted proteins. Rather than acting as a conventional enzyme inhibitor, a PROTAC works by inducing selective intracellular proteolysis. A heterobifunctional molecule with two active domains and a linker, PROTACs consist of two covalently linked protein-binding molecules: one capable of engaging an E3 ubiquitin ligase, and another that binds to a target protein. Recruitment of the E3 ligase to the target results in ubiquitination and subsequent degradation of the target protein via the proteasome.
It’s fun to read this sentence and now have a pretty good idea of what these things are! CRLs are examples of E3 ligases. That said, this is not really related to the challenge at hand. But it’s fun to make these connections and learn a bit more about a class of drug that of which I knew only the name.
Back to RBX1
Okay so last question before we get going - what kind of dysregulation is there in cancer? I would guess, if we’re targetting RBX1 with an antibody, that we want to prevent the formation of CRLs or the E3 ligase action of them. Which makes me think that in cancer, this system is degrading proteins that it’s not normally supposed to.
Looking into this, this paper gave me some basic understanding of why this is a good target. In short, turning off RBX1 with siRNA shrinks tumours! (Note that the paper calls RBX1 ROC1).
ROC1 silencing by siRNA significantly inhibited the growth of multiple human cancer cells via induction of senescence and apoptosis as well as G2/M arrest. […] Thus, ROC1 silencing triggers multiple death and growth arrest pathways to effectively suppress tumor cell growth, suggesting that ROC1 may serve as a potential anti-cancer target.
The paper is very much over my head but one thing that stood out in this paper and others I looked at to understand the relevance of RBX1 is that CRLs are a key part of regulating the cell cycle. Seems relevant to cancer! At any rate, the paper shows a clear therapeutic potential to the downregulation of RBX1, and that’s enough for me to know for now.
Cutting up RBX1
Okay so there seems to be good reason to target RBX1. And furthermore, we definitely want to affect its function as part of an E3 ligase in a CRL complex. So, we’re going to look at its role in CRLs, with CUL1 and CUL2 (although I think there’s 5 or 6 of these Cullinses) and try to understand more. The “how to make an antibody” guide suggests identifying a part of your protein that you want to target, and trimming the protein down to mostly that region to speed up the antibody generation process. So the next step was to figure out what part of the RBX1 protein we want to target.
Uniprot provides the following domains on RBX1:
- CUL1 binding: 5-108
- Zinc finger: 53-98
But uniprot also showed some crystal structures exist of RBX1 in complex with CUL2. I wanted to learn more about that and how the proteins bind, to inform what part of RBX1 we might want to target with our antibody. I was also confused by the overlap of CUL1 binding region and the Zinc finger. Almost the entire protein is present in the CUL1 binding region (RBX1 is 108 residues long), which seemed kind of curious to me.
That investigation lead me to this paper: Crystal Structure of the Cul2-Rbx1-EloBC-VHL Ubiquitin Ligase Complex which provided a crystal structure of RBX1 bound to CUL2. Here’s what they say:
Rbx1 structure presents an N-terminal tail arranged in a long β strand that engages Cullin CTD and a variant RING finger domain. The RING is characterized by the presence of an extended region containing a third zinc ion coordinated by three cysteines and one histidine, in addition to the canonical region containing two zinc ions (Zheng et al., 2002). The structure of the N-terminal tail of Rbx1 and its position in the complex are conserved among different CRL structures. Conversely, the RING domain is flexible and has been found in a number of different orientations, even in different molecules within the same ASU (Duda et al., 2008).
At first, this made me think we should target the N-terminal tail which binds to Cullin proteins. That way, we could prevent that binding to CUL1/2/etc. and avoid the formation of this complex which seems to be key to the re-regulating CRL-mediated protein degradation. However, looking at the protein in PyMol made me think otherwise. This is just my intuition (and it’s probably wrong) but, pointing an antibody at this long and generic protein strand seems like it would be a recipe for disaster. I don’t know for sure, but that just seems like it would produce a very low-specificity antibody.
So then I pivoted to focus on something more stable and possibly unique, which I decided would probably be the RING finger domain.
Leading off of that idea, I loaded up the crystal structure from the above paper in PyMol, and began to poke around.
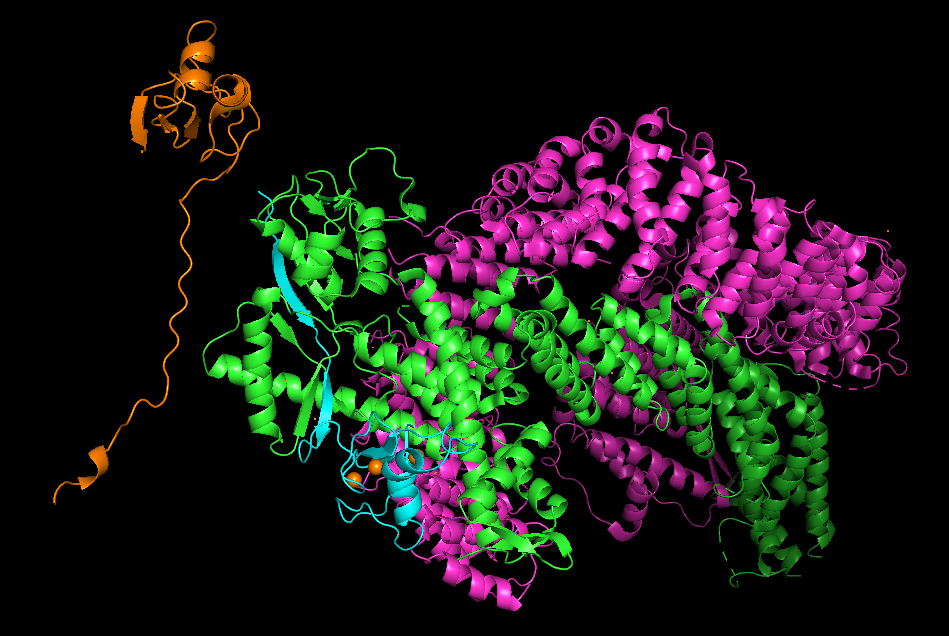
I then added in aligned the alphafold structure of RBX1 to the structure. Initially I did this because I was struggling to work out what part of the complex was RBX1. I had turned on per-chain colouring for the complex, which helped break it down but I wasn’t sure yet what was what. The alphafold RBX1 structure is in orange in the image above, and it turns out that RBX1 is cyan within the CRL complex.
I wondered if there might be a way to line up my two copies of RBX1, and it turns out, there is! I used the alignment plugin in PyMol with default parameters and it lined them up very nicely!
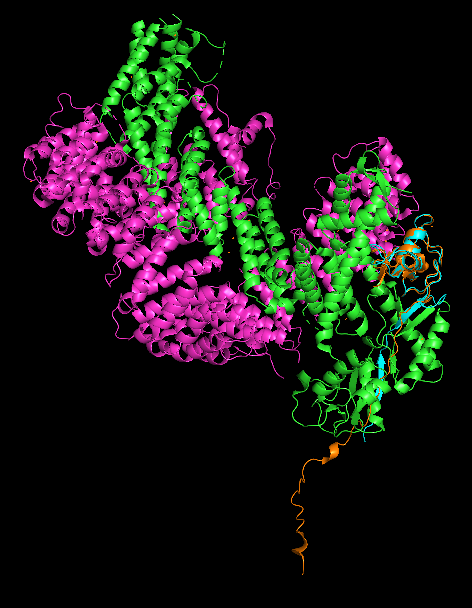
This gave us an interesting result - the N-terminal tail of Rbx1 is unconstrained in the AlphaFold structure and just sticks straight out, not aligning to what we see in complex with CUL2. The other end of the protein, with the zinc finger, is highly aligned however. So I turned on sequence mode in PyMOL and filtered to the region of our protein which was highly aligned - residues 36-108.
At this point, I was finally ready to make some antibodies! I exported my protein structure and then ran boltzgen in antibody-anything mode with default parameters to make 10 antibodies.
I pointed boltzgen at the zinc finger end of the trimmed protein as that’s what I think we want to engage.
I ran this on a paperspace instance to have GPU access, and it finished in a few minutes.
Result: I made an antibody?
I guess so? Let’s try AlphaFold and see what my antibody looks like in action!
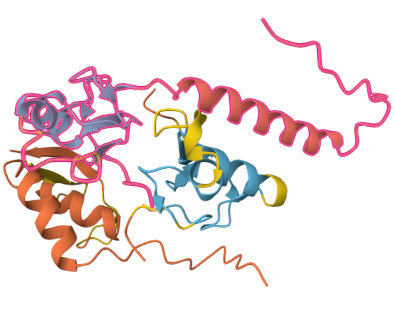
That looks… good? The highlighted protein is Rbx1. The proteins are colour-coded by AlphaFold’s confidence score in the position. It’s interesting that the RING finger area and the parts of the antibody that are closest to it are fairly high certainty! That seems good!
At this point of the experiment, I wanted to learn more about what I’d achieved, but my day was winding down just as this new door was opening. That will have to wait for another day.
Finally, I turned on the illustration mode in AlphaFold to get this pretty picture of my antibody in action:
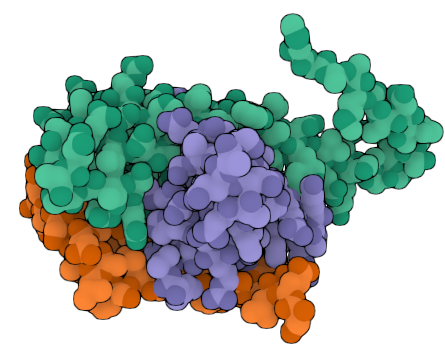
Open Questions
- What kind of antibody is this? I had assumed we were making a mAb but it doesn’t look like it!
- Alphafold wouldn’t have the zinc atoms in place in the RING finger domain as it takes just a FASTA sequence as input. Presumably this would affect binding?
- The antibody has two protein chains. One of them is very low certainty in Alphafold - is it doing anything?
- What’s the Kd for my antibody, is it any good? Would it prevent RBX1 from forming a CRL?
- Is there a safe/effective way/reason to target the N-terminal tail of Rbx1?
- did I just vibe-code an antibody?